01 · Measure
Stop reformatting cycler data. Start doing engineering.
Ionworks gives battery R&D teams structured data management for cycling data, cell metadata, and test measurements in one system of record.
Formats
·Maccor·Neware·Arbin·BioLogic·Novonix
Built by the team behind PyBaMM, the most widely adopted open-source battery modeling framework.
The structure problem in battery lab data
Most battery labs have plenty of data. The information exists, but it is scattered across formats, folders, and tools in ways that make it hard to use reliably.
01
Cycler data arrives in incompatible formats
A lab running Maccor and Neware simultaneously gets two different file formats with different column names, step conventions, and cycle definitions. Before anyone can compare results across test stations, someone writes a format-specific parser. Those parsers accumulate. They break when firmware updates change export schemas. The format problem never actually gets solved.
02
Metadata drifts away from measurements
Cell specifications, electrode compositions, electrolyte details, and formation parameters live in spreadsheets or lab notebooks that are updated on a different cadence than the cycling data they describe. Six months later, an engineer trying to reproduce a result has to reconstruct which cell was tested under which conditions. That reconstruction is where errors enter.
03
Analysis time goes to data plumbing
Every new analysis begins with file parsing, column renaming, and manual joins between test data and metadata. The team's Python scripts accumulate format-handling code that has nothing to do with battery science. Test engineers spend time on plumbing. Modelers wait for clean inputs. Two analysts looking at the same dataset may not be working from the same baseline.
What Ionworks provides
Multi-vendor cycler ingestion
The ionworksdata Python library reads raw files from Maccor, Neware, Arbin, BioLogic, and Novonix through named readers and normalizes them into a common time-series format. Computed columns for capacity, energy, and step tracking are added automatically.
Cell-centric data organization
Records are organized around Cell Specifications, Cell Instances, and Cell Measurements. Every measurement links to the cell it came from and the conditions it was tested under.
Connected data layers
Time-series data, step summaries, and cycle metrics are stored as connected layers within each measurement. A summary value can always be traced back to the underlying time-series record. Checkpoint measurements like cell images and thickness values sit in the same framework.
Programmatic and visual access
The Python SDK and REST API give analysis scripts and modeling pipelines direct access to structured data. Ionworks Studio provides a browser-based interface for upload, filtering, and visualization. Both work from the same measurement records.
How it works in Ionworks
01
Process and upload cycling data
The ionworksdata library processes raw cycler files into a common format through named readers (maccor, neware, biologic_mpt, and others). Time-series output includes automatically computed fields for step count, charge and discharge capacity, and charge and discharge energy. Original vendor-specific step and cycle values are preserved so nothing is lost during normalization.
Once processed, the Ionworks API handles upload. The CellMeasurementClient manages measurement data and supports scalable handling of large test datasets. Teams script the full ingestion pipeline in Python, making it repeatable across projects.
02
Structure measurements around cells
Ionworks organizes data in a hierarchy: Cell Specification, Cell Instance, Cell Measurement, with time-series data, step summaries, and cycle metrics accessible under each measurement record. Step summaries can be extracted directly from processed time-series data.
Beyond cycling data, Ionworks stores checkpoint measurements: non-time-series records captured at specific points in a test campaign, such as cell images, thermal camera output, or thickness measurements. All data associated with a cell is accessible through one system.
03
Visualize and review in Studio
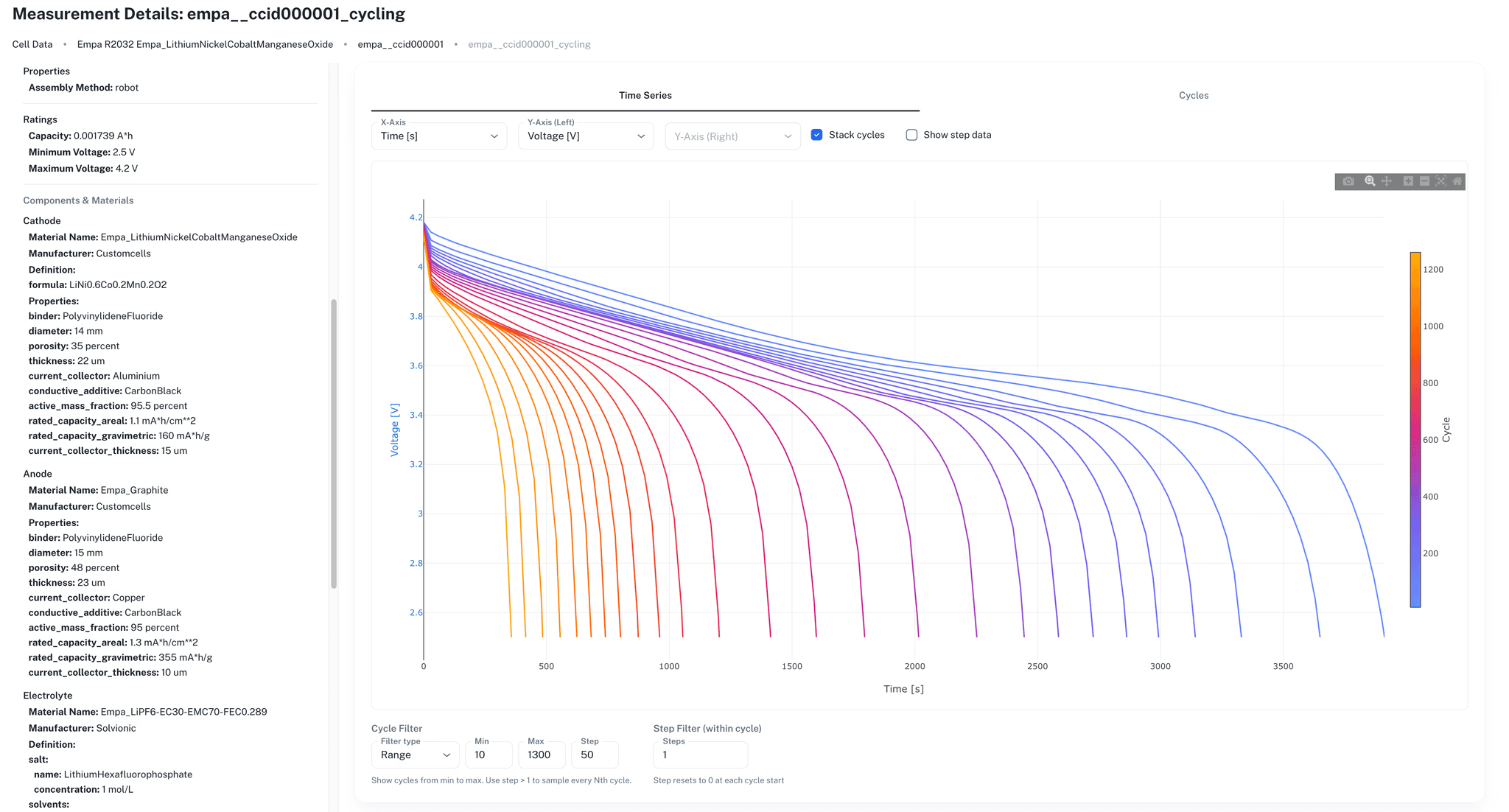
Ionworks Studio provides interactive tools for uploading, organizing, and visualizing cycling data in the browser. Engineers filter datasets, compare cycling curves, and inspect step or cycle summaries without writing code.
Studio is one access layer alongside the Python SDK and REST API. A test engineer might use Studio to spot-check a batch of new results while a modeler pulls the same data programmatically for a parameterization script.
04
Connect to model-driven workflows
Structured lab data is the starting point for battery model development. When time-series measurements, step summaries, and cell metadata are organized and accessible through an API, parameterization workflows pull real experimental inputs directly rather than re-parsing raw files.
Ionworks connects Measure to Train, Predict, and Optimize as part of the broader Simulation OS. For teams using PyBaMM, Ionworks provides the structured data infrastructure that modeling code assumes but does not provide on its own.
Frequently asked questions
Your cycler data is already scattered. Let’s fix that.
Import from Maccor, Neware, BioLogic, or Arbin and have structured, well-attributed test data in one session.