02 · Train
Stop guessing which parameter set is correct. Start tracking it.
Ionworks gives battery R&D teams repeatable model fitting and validation with full provenance, so every parameterized model links back to the data, the fitting process, and the cell specification that produced it.
Models
·SPM·SPMe·DFN·ECM·LumpedSPMR
Built by the team behind PyBaMM, the most widely adopted open-source battery modeling framework.
The parameterization bottleneck in battery modeling
Physics-based battery models depend on accurate parameterization, and many parameters are unknown or difficult to measure directly. The main issue is identifiability: with current as input and voltage as output, there is very little information to determine individual parameter values. Teams that model batteries already know the physics is the easy part. Getting the parameters right is where projects stall.
01
Parameter sets accumulate with unclear provenance
An engineer fits a model to a set of cycling experiments. The resulting parameters go into a spreadsheet, a Jupyter notebook, or a shared drive folder. Six months later, a colleague needs to reuse that parameter set for a new study. The notebook has been modified. The spreadsheet has multiple tabs from different fitting rounds. The original cycling data may have been reprocessed. Reconstructing which parameter set was validated against which data, under which conditions, with which fitting algorithm, becomes a project in itself. Teams end up refitting from scratch because trusting an inherited parameter set takes longer than creating a new one.
02
Identifiability limits what a single experiment can tell you
Each parameterization experiment is designed to amplify the contributions of a specific parameter subset while minimizing others. A GITT test constrains diffusion coefficients. A pulse resistance test pins down internal resistance. A slow-rate discharge reveals the open-circuit voltage curve. Combining different experiments determines the full parameter set. That makes parameterization time and resource intensive even when the fitting algorithm works well, because the experimental design matters as much as the optimization. Teams without a structured fitting pipeline often discover late that their data cannot constrain the parameters they care about.
03
Fitting workflows are hard to reproduce across a team
A parameterization pipeline involves choices at every step: which experimental data to include, which parameters to fix versus fit, which bounds and initial guesses to use, which optimization algorithm to run. Those choices live in a single engineer's notebook. When another team member needs to fit the same model type against a different cell, they start from a blank notebook. The institutional knowledge about which fitting configurations work, which parameter combinations are identifiable, which bounds prevent nonphysical results, stays with the person who did the last fit.
What Ionworks provides
Parameterized models as first-class objects
A parameterized model bundles three things: a specific electrochemical model type (SPM, SPMe, DFN, ECM, or LumpedSPMR variants), a validated set of parameters, and a cell specification. Once used in a simulation, a parameterized model becomes immutable. To modify it, clone it, creating a new version with clear lineage back to the original. Every parameterized model in the system links to the fitting process and the data that produced it.
Multiple fitting objective types
The ionworkspipeline package supports fitting against current-driven cycling, cycle aging, EIS, pulse resistance, OCP half-cell, electrode balancing, and MSMR (multi-site multi-reaction) data. Different experiments constrain different parameters. Combining objective types in a single pipeline builds a complete parameter set with each parameter constrained by the experiment designed to reveal it.
Sensitivity analysis and identifiability checks
Sobol sensitivity analysis quantifies which parameters actually influence the model outputs for a given experimental protocol. Before committing to a long fitting run, teams can check whether their data can constrain the parameters they care about. This surfaces identifiability problems early, before the optimizer runs for hours and returns a result that looks good numerically but has traded off between correlated parameters.
Six optimization algorithms with configurable starts
Fitting uses Differential Evolution, PSO, CMA-ES, XNES, SNES, or Nelder-Mead, with Latin Hypercube Sampling for initial populations. Population-based optimizers distribute candidate evaluations across workers. Multistart runs the optimizer from multiple starting points to reduce the risk of landing in a local minimum.
How it works in Ionworks
01
Select a model type and cell specification
Start with the electrochemical model that fits the engineering question. SPM is fast and suitable for many applications. SPMe adds electrolyte dynamics. DFN is the full-fidelity porous electrode model. ECM with configurable RC pairs (0-5) fits impedance and pulse data. LumpedSPMR and LumpedSPMeR add lumped resistance terms. All physics-based models support full-cell and half-cell configurations with isothermal, lumped, or two-state thermal coupling.
Link the model to a cell specification that defines the cell's materials, capacity, voltage limits, and form factor. The cell specification is shared across the organization, so teams fitting models against different batches of the same cell type start from the same baseline.
02
Configure parameters and run the fitting pipeline
Parameters take several forms: scalar values, expressions that reference other parameters, temperature-dependent functions (Arrhenius expressions), and interpolants (lookup tables for OCP curves, diffusivity, and other data-driven relationships). Interpolants support 1D and multi-dimensional inputs with linear, cubic, or pchip interpolation. Define which parameters to fit, which to fix, and set bounds that prevent nonphysical results.
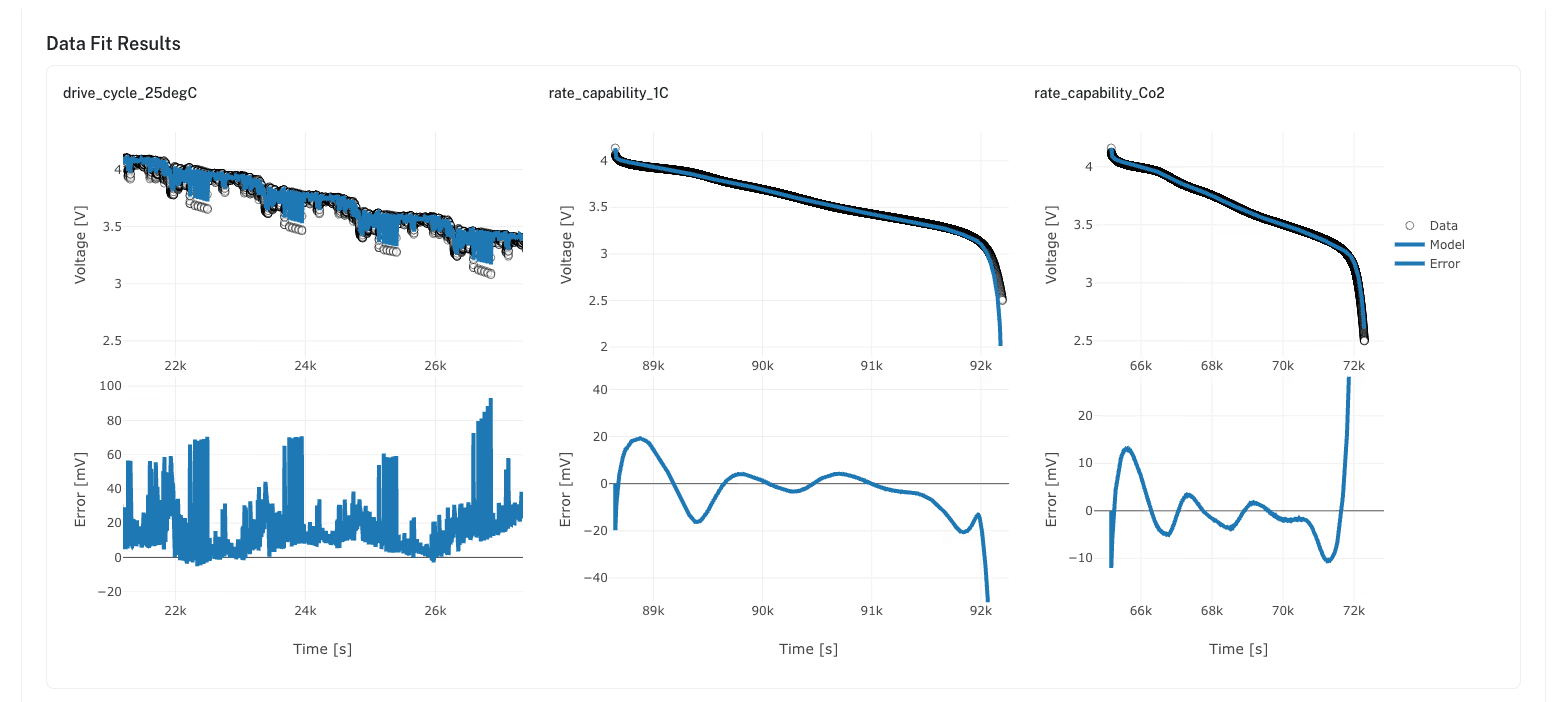
The ionworkspipeline package handles the full fitting workflow. Combine multiple objective types in a single pipeline: fit OCP from half-cell data, constrain kinetic parameters from pulse tests, validate against full cycling data. The pipeline runs as a background job with real-time status updates. Results include parameter values, voltage-vs-data comparison plots, and RMSE per objective.
03
Validate and version the parameterized model
Review fitting results against the experimental data. Compare the model's voltage, current, and temperature predictions against measured values across the protocols used for fitting and against held-out validation data. Custom variables let teams define derived quantities from model variables: electrode potentials, scaled parameters, power output.
Once validated, the parameterized model is ready for simulation. The first time it is used in a Predict or Optimize workflow, it becomes immutable. All subsequent modifications require cloning, which creates a new version with lineage back to the original. The full fitting history, the data used, the algorithm settings, and the resulting parameters are preserved as provenance.
04
Connect to simulation and optimization workflows
A validated parameterized model feeds directly into the Predict and Optimize stages of the Simulation OS. Engineers select a parameterized model by name and version, define a protocol, and run simulations. The model's provenance travels with every result: anyone reviewing a simulation can trace back to the exact parameter set, the fitting process, and the experimental data that grounded it.
Teams that fit models against new cell batches clone existing parameterized models and refit against new data, building a library of validated models organized by cell specification and chemistry. The ECM parameterization tool provides a browser-based interface for fitting OCV, R0, and RC-pair parameters directly from cycling data, accessible to team members who do not work in Python.
Frequently asked questions
Bring a dataset. Leave with a parameterized model.
Walk through the full fitting and validation workflow with our team on your own data.