Take PyBaMM battery simulation from notebooks to team workflows
Ionworks gives battery R&D teams the data infrastructure and simulation management to turn open-source PyBaMM models into repeatable engineering processes.
PyBaMM alone vs. PyBaMM with Ionworks
PyBaMM alone
- Vendor-specific data parsing
- Parameters in notebooks and spreadsheets
- Manual simulation coordination
- No shared model versioning
- Results scattered across output folders
With Ionworks
- Common format, all major cyclers
- Parameterized models with full provenance
- Managed simulation runs, automatic reuse
- Immutable, versioned, shared across the team
- Studies with linked, traceable results
The real problem
Where notebook-based PyBaMM workflows break down
01
Battery parameter estimation is the real bottleneck
Physics-based battery models depend on accurate parameterization, and many parameters are unknown or difficult to measure directly. The main issue is identifiability: with current as input and voltage as output, there is very little information to determine individual parameter values. Over time, teams accumulate parameter sets with unclear provenance. Multiple engineers fit models to different datasets, store results in different notebooks, and keep no shared record of which parameter set was validated against which test data.
02
Test data is fragmented across lab systems
Battery R&D teams generate data across BioLogic, Maccor, Neware, Arbin, and other cycler platforms. Each produces its own file formats with different column names, step conventions, and cycle definitions. An engineer trying to reproduce a parameterization result six months later has to reconstruct which cell was tested under which conditions, which data files correspond to which experiments, and which version of a parsing script produced the clean dataset.
03
Scaling simulations past what notebooks can manage
Running a single simulation in a Jupyter notebook is straightforward. Running a structured study that compares dozens of protocol variations across multiple cell designs, tracks which parameterized model was used for each run, and lets a colleague reproduce results six months later requires infrastructure notebooks were never designed to provide.
04
Python flexibility becomes a coordination cost
When every engineer writes their own simulation scripts, each with different conventions for handling parameters, protocols, and results, the team accumulates coordination debt. A weekly simulation that three people run produces three slightly different workflows.
How Ionworks extends PyBaMM for teams
Ionworks Studio is a battery modeling platform built on PyBaMM. It adds data management, parameterization tracking, and simulation coordination so models become shared, reproducible engineering processes.
01
Measure
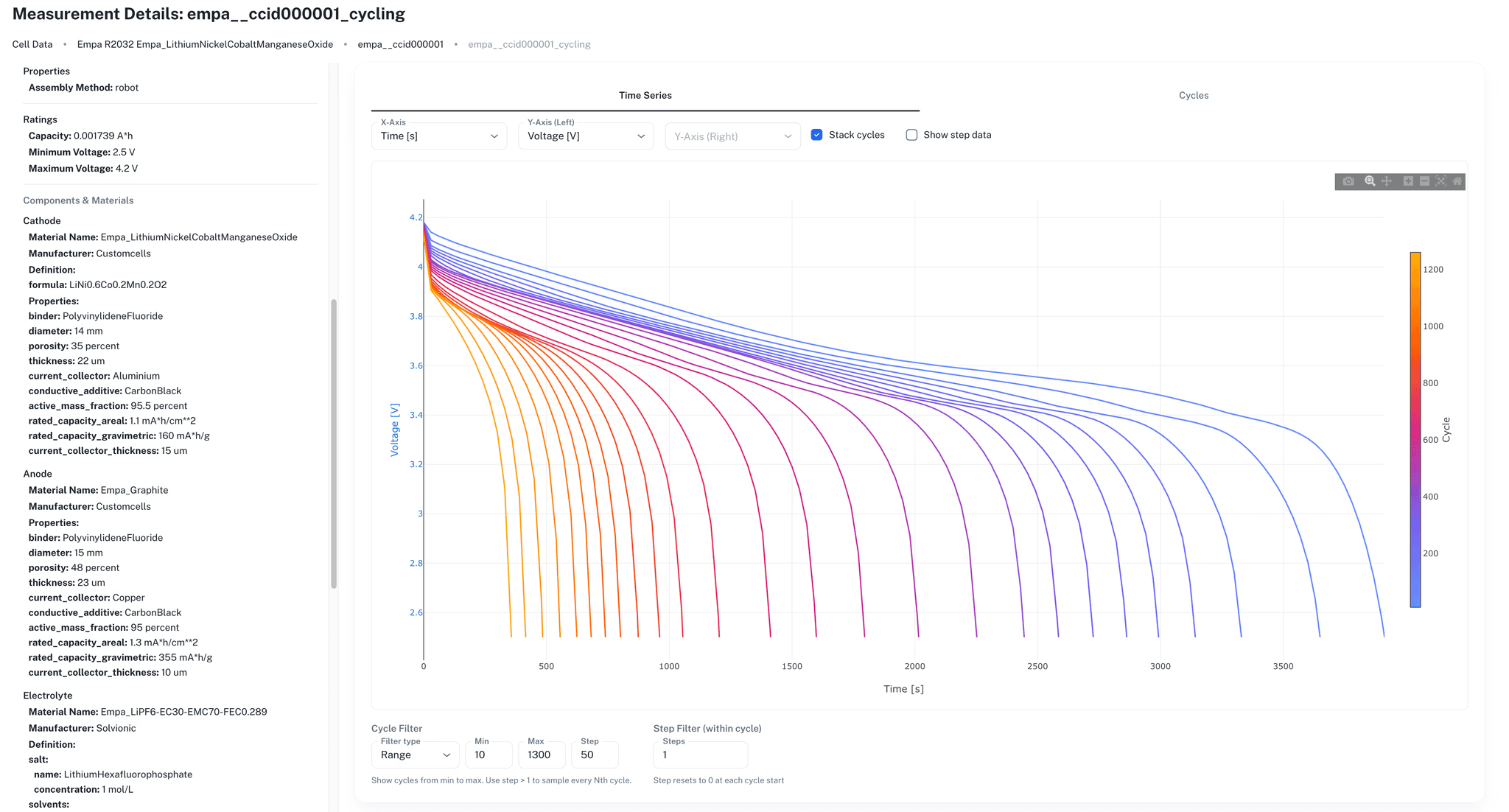
Upload data from Maccor, Neware, BioLogic, Arbin, Novonix, and other cycler formats into one shared system of record. The ionworksdata Python library processes raw files through named readers and normalizes them into a common time-series format with automatically computed fields for capacity, energy, and step tracking.
Data is organized around cell specifications, cell instances, and cell measurements. Every measurement links to its cell and experimental context. Teams script the full ingestion pipeline in Python, making it repeatable across projects.
02
Train
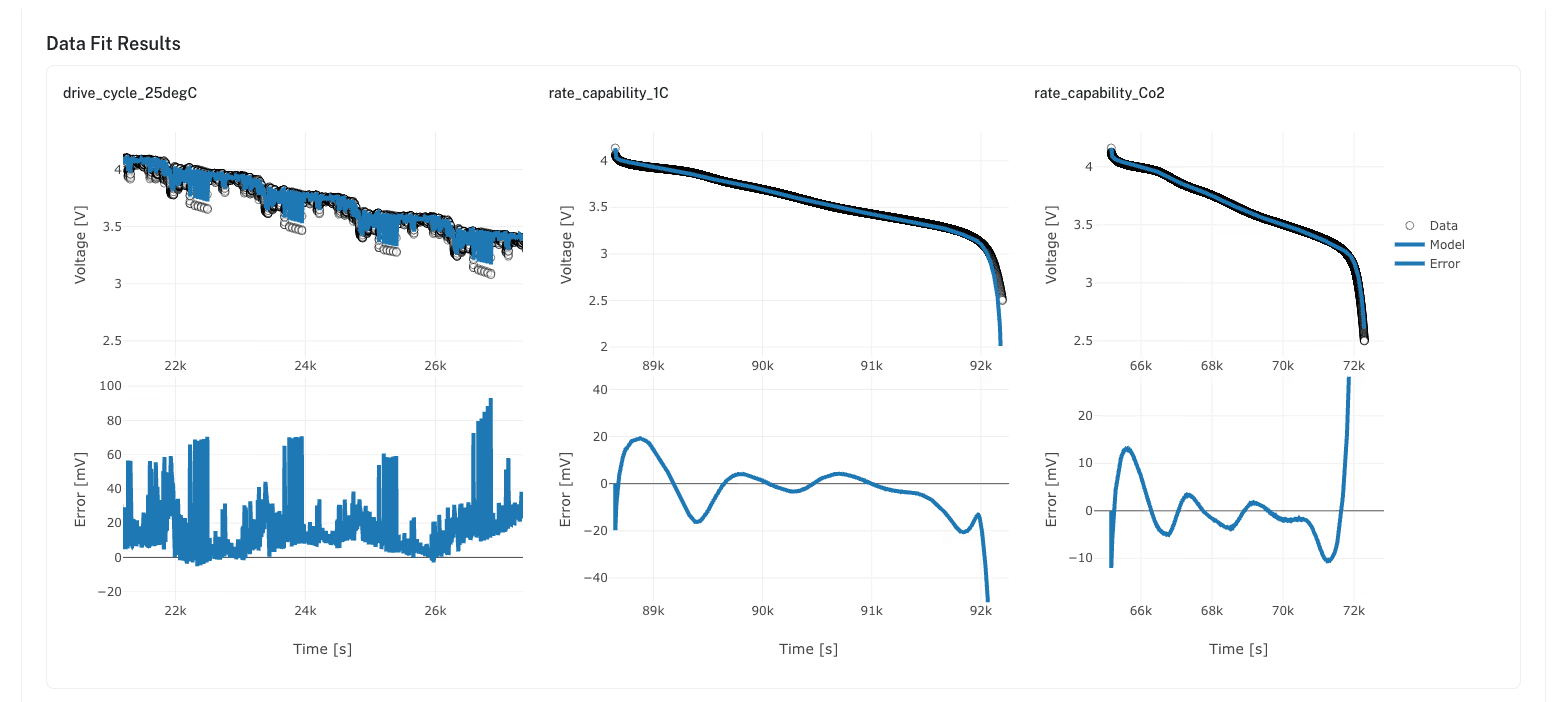
Build parameterized models through a visual modeling pipeline. Each parameterized model bundles a specific electrochemical model type (SPM, SPMe, DFN), a validated parameter set, and a cell specification into a single, ready-to-run configuration.
Once used in a simulation, a parameterized model becomes immutable. To modify it, clone it, creating a new version with clear lineage back to the original. No more spreadsheets of parameter values with uncertain origins.
03
Predict
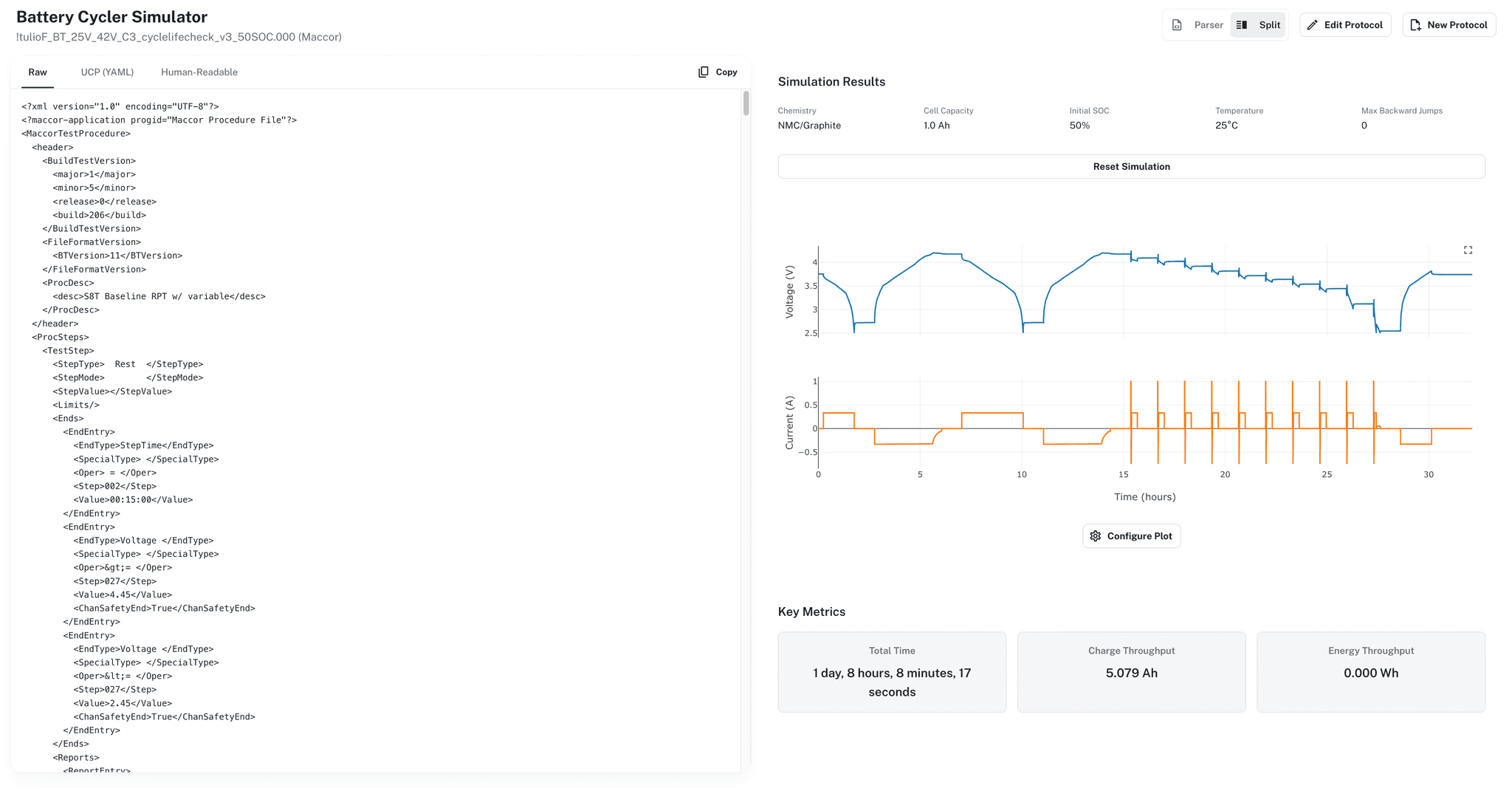
Select a parameterized model, define an experiment protocol, run the simulation. Protocols can be saved, typed as text, uploaded from cycler files, or built with the protocol builder. Studies group related simulations into focused investigations.
Identical completed simulations are reused automatically rather than rerun. The same parameterized model and protocol always produce the same result, and that result is stored once.
04
Optimize
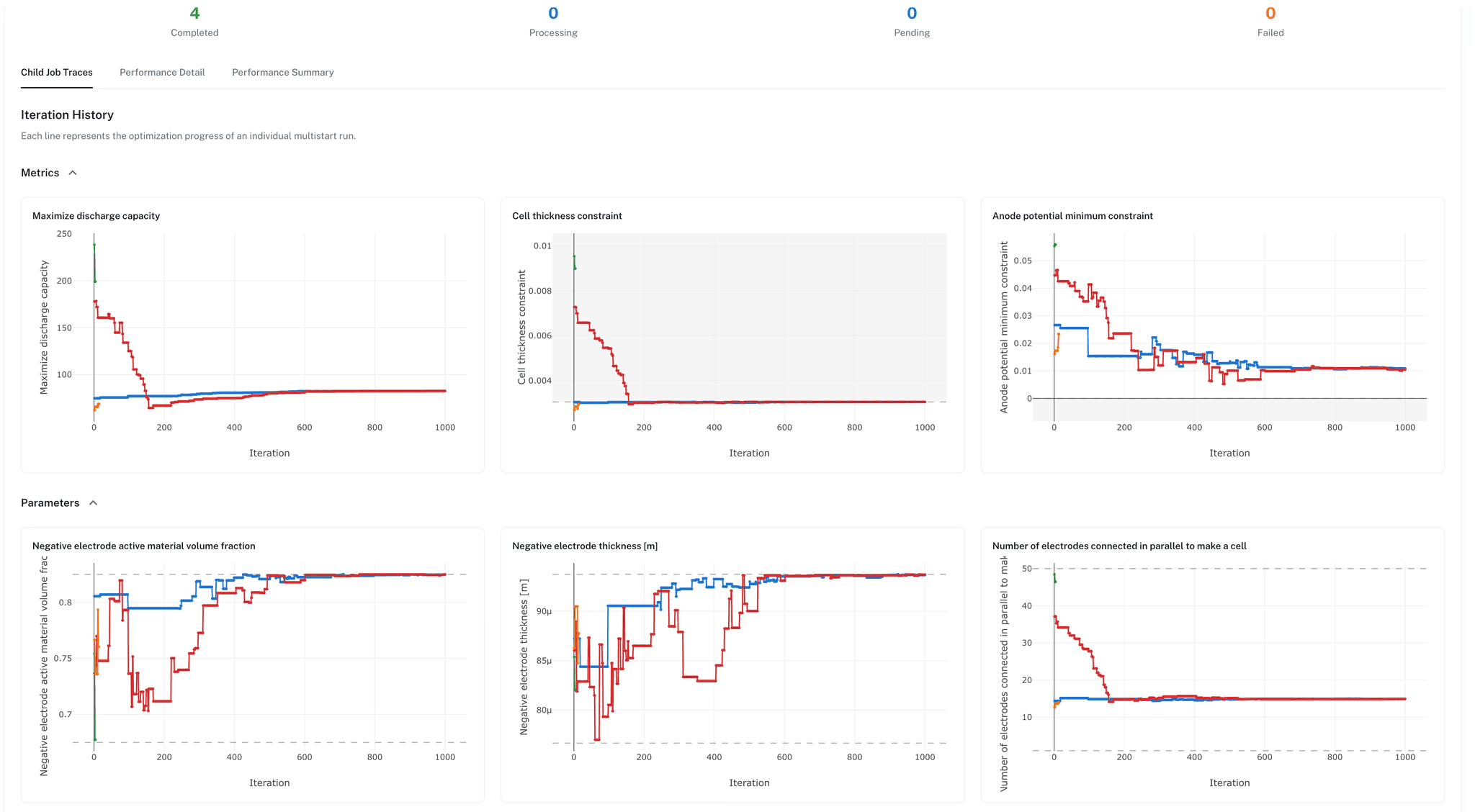
Sweep coating thickness, porosity, loading, and electrode geometry against real performance targets. Define objective functions tied to energy density, cycle life, charge time, or thermal headroom. Set constraints for voltage limits, temperature bounds, and plating thresholds.
The same parameterized models validated against test data now explore the design space. A design sweep that would take weeks of physical testing completes in minutes.
Example questions teams can answer
What is the fastest charge rate that avoids lithium plating across a temperature range?
A validated electrochemical model predicts plating onset across conditions that would take months to test exhaustively. Sweep C-rates and temperatures against plating thresholds in a single study.
How do we set charge and discharge limits for a new cell chemistry?
Parameterized models fitted to early test data let teams simulate operating envelopes before committing to long cycling studies. Define voltage, current, and temperature bounds from simulation rather than trial and error.
What are the energy-power tradeoffs for different electrode designs?
Sweeping design parameters in Ionworks Studio surfaces tradeoffs across the design space in hours. Vary coating thickness, porosity, and loading to map energy density against rate capability.
Which charge protocol minimizes degradation for a target duty cycle?
Protocol-driven simulations using real cycler files let teams compare protocol variants against the same validated model. Run degradation predictions across CC-CV, multi-step, and pulsed protocols.
Frequently asked questions

Ionworks enables our R&D Services customers with tools and insight that support faster development and more predictable outcomes.

Your PyBaMM models already work. Make the workflow around them work too.
If your team spends more time managing scripts and parameter files than interpreting results, Ionworks Studio is worth 30 minutes. Bring your own notebooks and we'll show you the difference.