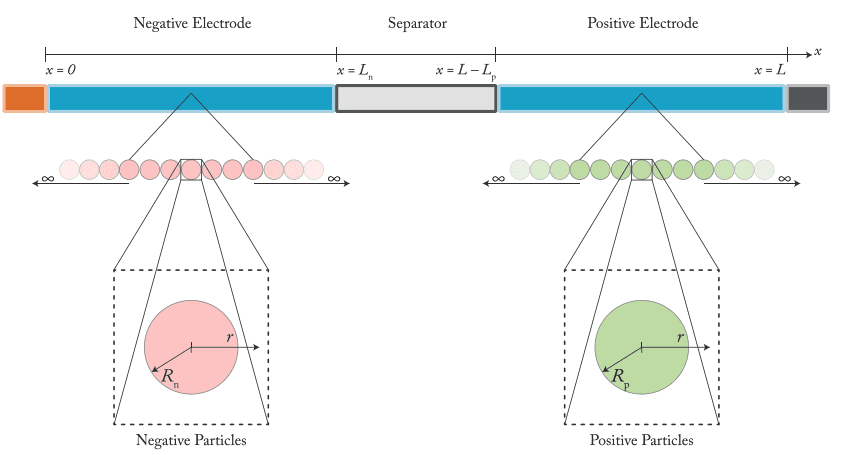
One of the main challenges in physics-based modelling of batteries, as discussed in our previous blog post, is parameterisation. By parameterisation we mean the process of determining the values of the model parameters so they accurately describe a certain situation, in this case, the cycling of a specific battery. For physics-based models, given the large number of parameters required (over 30 for the DFN model) and the fact that some of them are functions of the concentrations or temperature, parameterisation requires a broad range of experiments to be run. and the resulting data to be carefully analysed. This makes parameterisation an extremely time and resource intensive process. But why is that?
The identifiability problem
The main issue is the identifiability of the system. We say a system is identifiable if it is possible to infer its unknown parameters by measuring its output over time. In battery systems, especially in the parameterisation experiments, we typically have current as input and voltage as output (or viceversa). This means we usually have very little information to determine the parameter values, and thus the need for all the different experiments. Each experiment is designed to amplify the contributions of the parameter subset we are aiming to determine, while minimising the contributions of the remaining parameters. By combining different experiments, we can determine the entire parameter set for the model.

Separating the contributions of each electrode
A particular problem is separating the contribution of each electrode, hence the need for half-cells or three-electrode cells. A good way to illustrate this issue is to consider a very simple electrical system: a voltage source with two resistors in series (see Figure 1). Imagine the voltage source is set to 2 V and we measure the current to be 1 A. In this situation, we know that the total effective resistance of the circuit is 2 Ω, but we have no way of knowing what’s the actual value of each resistance, it could be R_1=R_2=1 Ω, or R_1=0.5 Ω and R_2=1.5 Ω, or any other combination of values: they would all predict the same result. The same occurs when we measure the voltage of a battery: the contributions of each electrode superpose and it is not possible to identify the parameter values for each electrode. However, going back to our simple example, if we could get some additional measurement, like the voltage drop across one resistor in our example circuit V_meas, then we would be able to uniquely determine the value of each resistance. Similarly, if we can measure the potential of each electrode against a reference electrode, we can determine the parameter values for each electrode.
Noise, variability, and what Ionworks does about it
Telling apart each electrode is only part of the identifiability problem. In real applications, handling the noise in the measurements or cell-to-cell variations add even more complexity to the parameterisation process. Ionworks builds on PyBaMM to make this process tractable. PyBaMM’s flexibility lets us compare multiple models and pick the right one for each application. Its fast solvers let us fit the chosen model to match experimental data.
Frequently asked questions
Continue reading